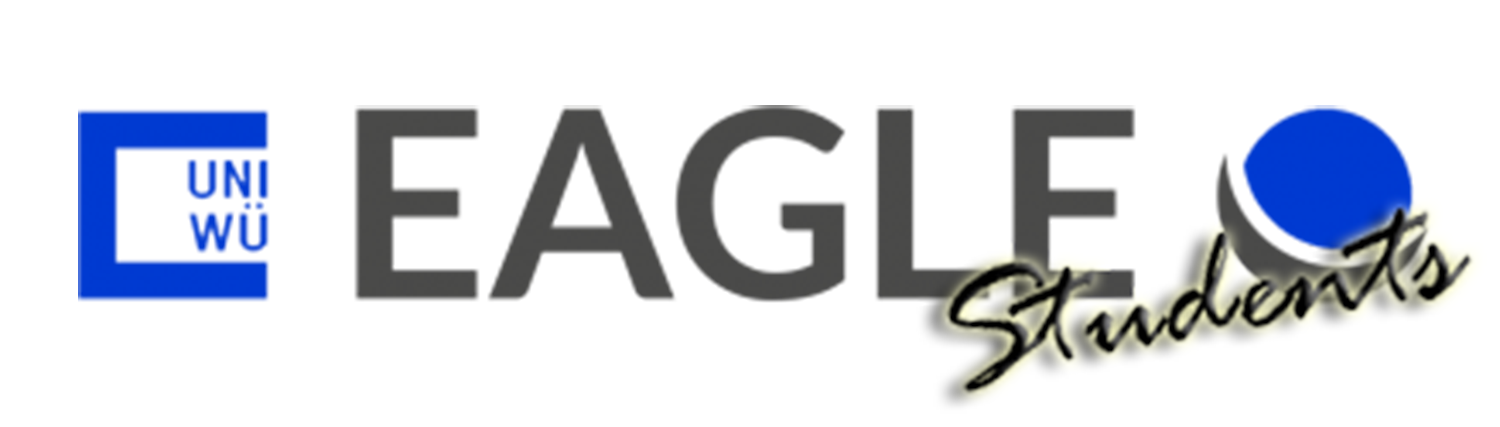bubblegam R package
For our "Programming & Statistics for Remote Sensing and GIS" course we have developed an R package to efficiently merge (geo)dataframes, identify, and move "spatial outliers" in geodata, create geographic plots, bubbleplots, and the animation between these plots....
EAGLE Clara at EGU Snow Science Winter School in Finland
Hey, it’s me, Clara! Last week I was able to participate in a winter school on snow field methods and microwave modeling of snow and wanted to share some impressions here. Using bus and train I traveled to the research center of the Finnish Meteorological Institute...
8th generation EAGLEs – First semester recap
Our first semester is ending, and a lot has happened in the last few months since classes started, so it's time for a little recap! In mid-October, we were welcomed in the seminar room at John-Skilton-Straße 4 on the northeastern edge of the campus. Thus, we were the...
Social Event at Boulder Hall
Last Friday, students and staff at the Department of Earth Observation Research traded the lecture rooms for a bouldering hall where they came together for a social event. With feet squeezed into climbing shoes and chalky hands, the participants embraced the challenge...
8th generation EAGLEs participate in Mapathon
The eighth generation of EAGLE students got off to a good start in their first semester! This Tuesday evening, following the afternoon session of "Introduction to Programming and Geostatistics", some EAGLEs participated in a Mapathon. It was organized as part of the...
EAGLEs at DLR
After finishing their first exam, the young eagles of the 7th generation travelled on the 8th of December to Munich to celebrate it at the Tollwood/Christmas Market. On Friday, they were invited to visit the DLR-EOC in Oberpfaffenhofen to get an insight into different...
ESA LPS 2022- EAGLES experiencing the largest EO conference!!
It was a huge week filled with talks, posters, and lots of networking sessions for the EAGLES at ESA Living Planet Symposium 2022. The whole week was arranged for the symposium with no courses taking place, which showed how excited the EAGLE staff and students were...
Eagles in Winter Wonderland – a trip to Berchtesgaden
After 2 months of back-to-back courses, one exam, and a lot of studies the EAGLE of the 6th generation took a break at Berchtesgaden enjoying a lot of snow!!The group of 14 left for this winter wonderland on December 9th after an informative seminar session on GDAL....
Either EAGLE or Ibis?
Studying remote sensing is important to take one step back being able to see changes in a whole, because we see them from space. For me, a 4th Gen EAGLE, sometimes the analyses are not enough. This is why I would like to share my story with you, about what I am doing...
EAGLE students stretching their Wings – Paragliding Trial Course
On the weekend of 17th and 18th of October, three EAGLEs met at the “Wasserkuppe”, the highest mountain of Hessen in the landscape called “Rhön”. Daily handling of airborne images as remote sensing student definitely makes you wishing to experience this bird view...
Review: First semester of the 4th Eagle Generation
The 4th EAGLE generation is already slowly approaching the end of the first semester and already a lot has happened in this first months. One highlight was visiting the DLR's Earth Observation Center (DLR-EOC) in Oberpfaffenhofen close to Munich in December. There...
Impressions from the EAGLE students’ participation in the 2019 Geo Summer Party’s Dodgeball Competition
Instead of hosting a football competition as it always had been for the last years, the Geo Summer Party 2019 Planning Committee instead decided to introduce a new game: Dodgeball. As last year, EAGLE students formed a team representing the Department of Remote Sensing at the University of Würzburg and fought hard to survive the competition. Browse through the images below to get some impressions of the event 🙂
EAGLEs @ ESA Living Planet Symposium 2019
From May 13 to 17 the city of Milan hosted the world’s largest Earth Observation conference, the ESA Living Planet Symposium. Over 4000 participants (remote sensing experts, national and international companies, students, organizations and agencies from all over the...
EAGLES at ‘Big Data from Space’ Conference
Last month, from 19-21 February 2019, a group of EAGLE students participated at the “Big Data from Space” conference in Munich. The conference co-organized by ESA & DLR focused on three main topics: 1) Latest advances in machine and deep learning 2) Latest...
First EAGLE generation: Spreading its wings and taking off
Since more and more EAGLEs of the 1st generation started to stretch their wings, it is time for one last recap. Over the last two and a half years, every one of us was quite busy learning new and exciting techniques and methods in the scientific field of remote sensing. We also were given the chance to expand our network and explore academic fields, which were not necessarily related to our previous occupations.
The EAGLES on the road at DLR and in the Alps
Last week, on December 7, the latest EAGLE generation also made its way to Oberpfaffenhofen near Munich to visit DLR’s Earth Observation Center (DLR-EOC). This year there were again very exciting talks on the following topics: Optical radar remote sensing for...
The 2018 EAGLEs are online!
[code language=”R”]print(“Hello, World!”)[/code]
Today, the new EAGLE intake that pooled from all over the World has completed six academic weeks in the program – it’s about time to introduce ourselves!
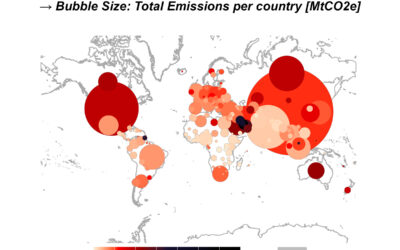
Settling down in Würzburg – an International’s edition
Settling down in a foreign country can be a challenging process, especially if you do not speak their language well. Untangling the process can be daunting, complex, and usually lengthy.
EAGLEs forming a football team at yesterday’s Geo summer party
Every year, many Geography students and some lecturers of the University of Wuerzburg meet at the center lawn of Campus Hubland for the annual Geography summer party, organized by the faculty’s Student Council. This year, as every year, one highlight was the football tournament, in which nine teams, all drafted explicitly for the event, competed against each other to win the “Mohorovičić Cup”.
Summer Dialogue 2018
We will be holding our annual EAGLE Summer Dialogue and Barbeque on 22 June 2018. Here is a quick peek at our programme:
Field Measurements in Demmin 2018
In order to get some practical experience in terms of field work the EAGLE program offers an excursion to Demmin which is located in federal state of Mecklenburg-Western Pomerania in North-East Germany. The test site is an intensively used agricultural ecosystem that...
Block course “Hyperspectral Remote Sensing”
One reason that makes this master’s program so attractive is its highly diverse coursework. This is being reflected by the currently available block courses. The “Hyperspectral Remote Sensing” course held by Dr.rer.nat. Martin Bachmann is part of that. In this course...
Photoshooting for the EAGLE Website
Another Year, another generation of students. After our studies began there was also the need for updating the EAGLE students’ webpage in order to reveal the new generation of EAGLEs to the world. The first step was of course the creation of a group photo, as well as...
2017 EAGLE students’ website now online
The 2017 EAGLE students' website is now online, featuring portraits of the students including their background and special interests.
Bavarian Mountains, East-German Landscapes and a Party: The First EAGLE Summer Term
Now the new EAGLEs are arriving and an exciting second year is about to start. But first let’s take a look back at an enjoyable summer term. We were right off to a promising start by visiting the KIT (Karlsruhe Institute of Technology) Campus at Garmisch-Partenkirchen...
students are currently studying in the EAGLE M.Sc. program
students have already graduated from EAGLE
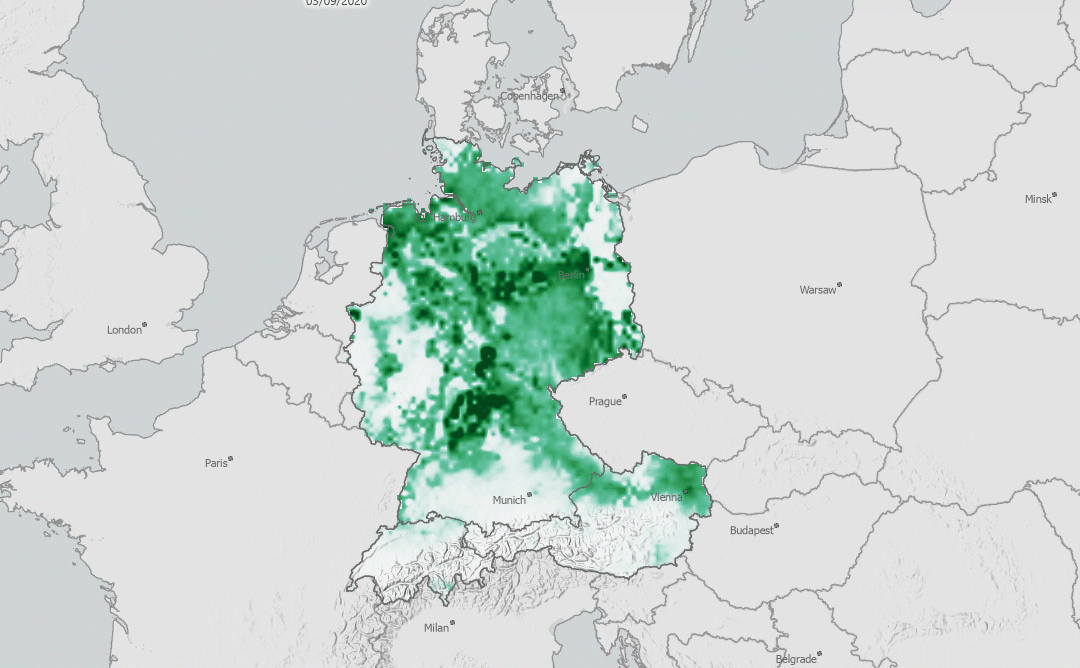
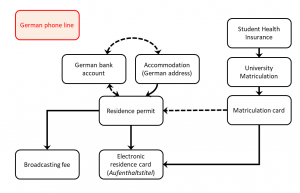
Processing of Sentinel-5P data: Creating nitrogen dioxide time series in R and ArcGIS
written by Magdalena Halbgewachs Sentinel-5P is a satellite that is operating within ESA's Copernicus program since 2017. The goal of the mission is a very dense scheduled operational monitoring of the atmosphere. Using the TROPOMI instrument on board, various...
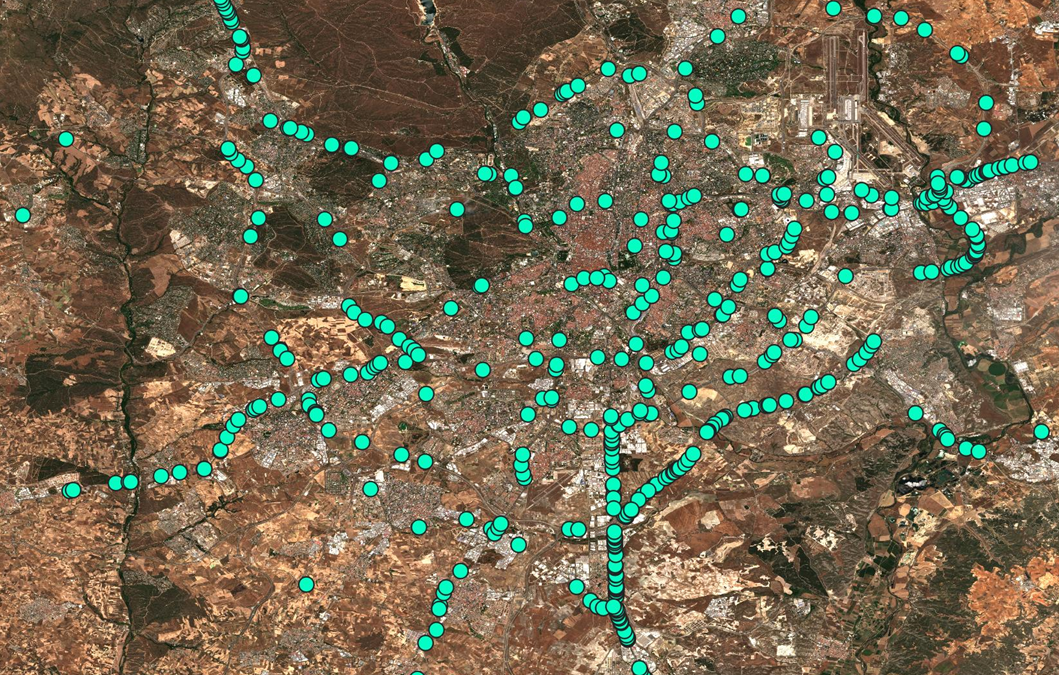
Truck Detection using Sentinel-2 data
In the context of the COVID-19 Custom Script Contest organized by ESA and Euro Data Cube Henrik Fisser developed a Sentinel-2 truck detection method. It relies on an effect that spectrally exposes moving objects of a certain size in Sentinel-2 data. This effect is caused by the layout of its Multispectral Instrument (MSI). In the following article he will give you an explanation of the effect, the method development and his experience upscaling the detection to whole EU extent for integration into the ESA RACE dashboard.
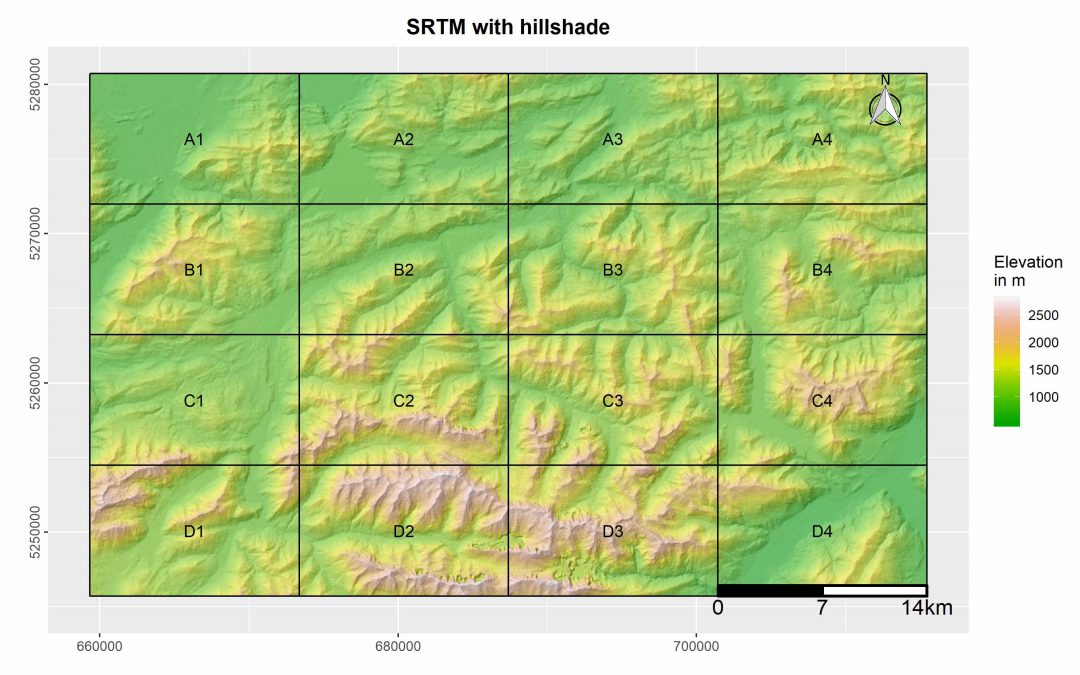
R package “MapBooklet” for an automatized creation of map booklets based on user ggplots
This R package was designed by EAGLE student Marius Philipp for the automatized generation of map booklet pdfs for user defined ggplots. The package contains currently two functions, one of which creates a fishnet polygon from the extent of the user input data. The second function uses these fishnet polygon in combination with a user ggplot for the creation of a map booklet, including an overview map, as well as a submap for each tile.
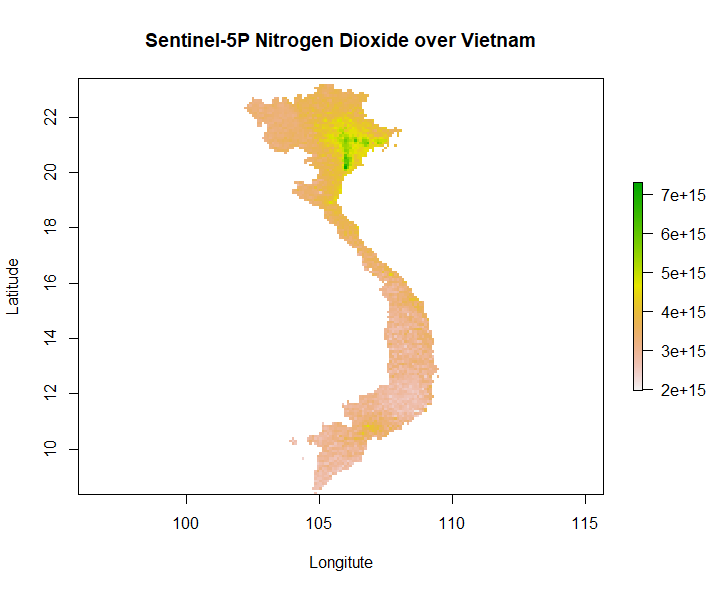
R Processing Toolbox for Sentinel-5P data
Our second generation EAGLE student Marius Philipp developed the R package S5Processor to automatically convert Sentinel-5P L2 NetCDF files into TIFF files. The user can define the product which should be turned into a TIFF and also mosaic from a list of file-paths....

An EAGLE student’s contribution to a study on mapping farming systems in Africa to be presented at ESA Living Planet Symposium 2019
In a few days, me and some other EAGLE program students are going to attend the “Living Planet Symposium 2019” by ESA which is amongst the biggest Earth observation conferences in the world. Part of my internship work at Remote Sensing Solutions GmbH is going to be...

Salim Soltani’s internship experience at DLR site Oberpfaffenhofen
The internship at DLR gave me the opportunity to contribute to a case study investigating the effect of spatial resolution on the snow classification accuracy using different sensor images using Spot-5, Sentinel, Landsat, and MODIS for the time frame of 1984 to 2018....
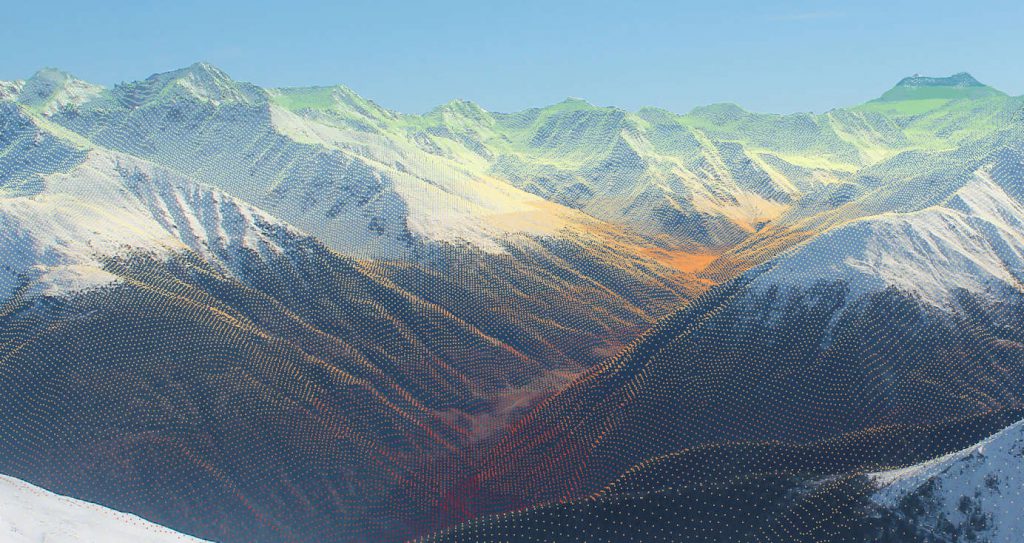

georef_webcam – A python toolbox to georeference webcam images
Over the last years, webcams and real-time monitoring has become more and more popular. Especially in mountainous regions webcams are widespread (e.g. ski fields, lakes, traffic, …). As these images provide information in high spatial and high temporal resolution, the...
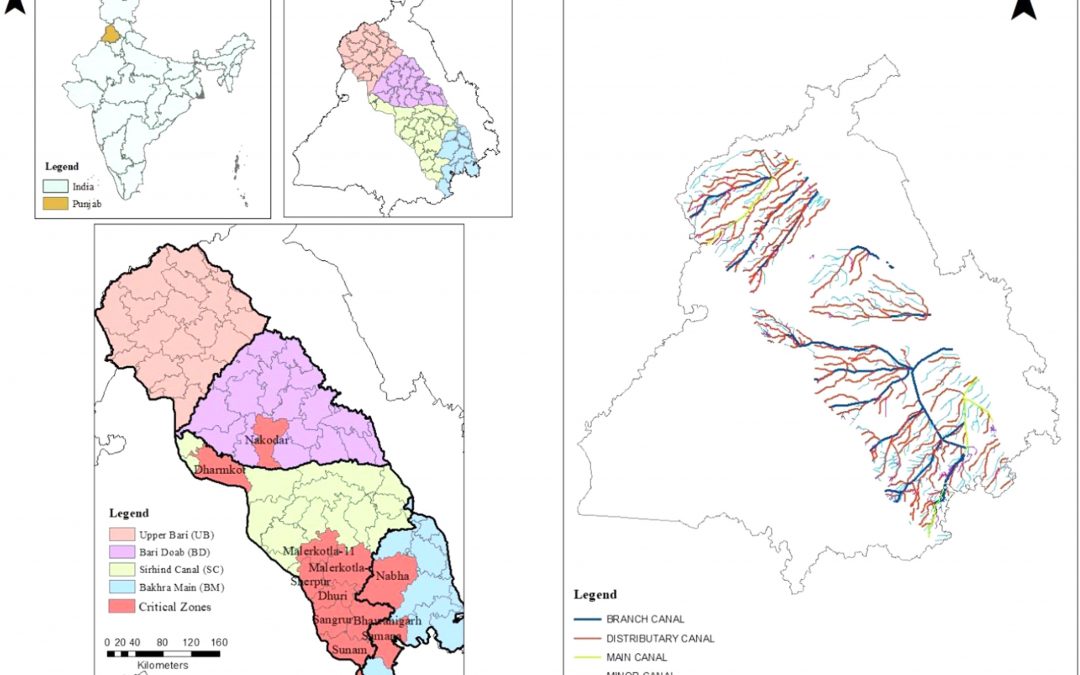
Methodology to delineate the critical regions for mitigation of carbon emissions due to groundwater pumping
Figure: Map of the study area and canal network of central Punjab with critical regions After estimating the total carbon emissions from groundwater pumping (see the previous blog), I wanted to delineate the critical regions of the study region with high carbon...
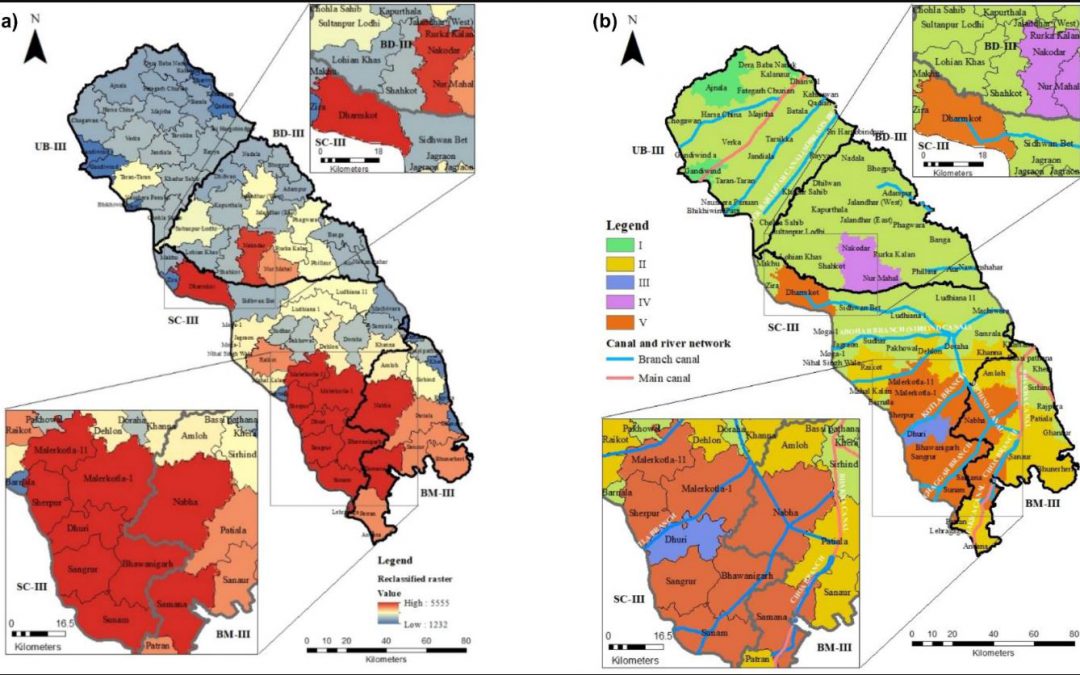
A methodology to estimate carbon emissions from groundwater pumping
Recently, I published a case report, which was based on a methodology to estimate carbon emissions from groundwater pumping in Punjab, India. The report was correlated with the fact of the green revolution momentum in the late 1960s and 1970s that resulted very...

Two 2016 EAGLE students win ESA Copernicus Hackaton 2018 in Darmstadt
Written by Karsten Wiertz This year, two of our EAGLE students had the opportunity to participate at the Copernicus Hackathon 2018 in Darmstadt. In the Hackathon, participants formed small groups, chose a task and had 24 hours to work on it with the technical support...
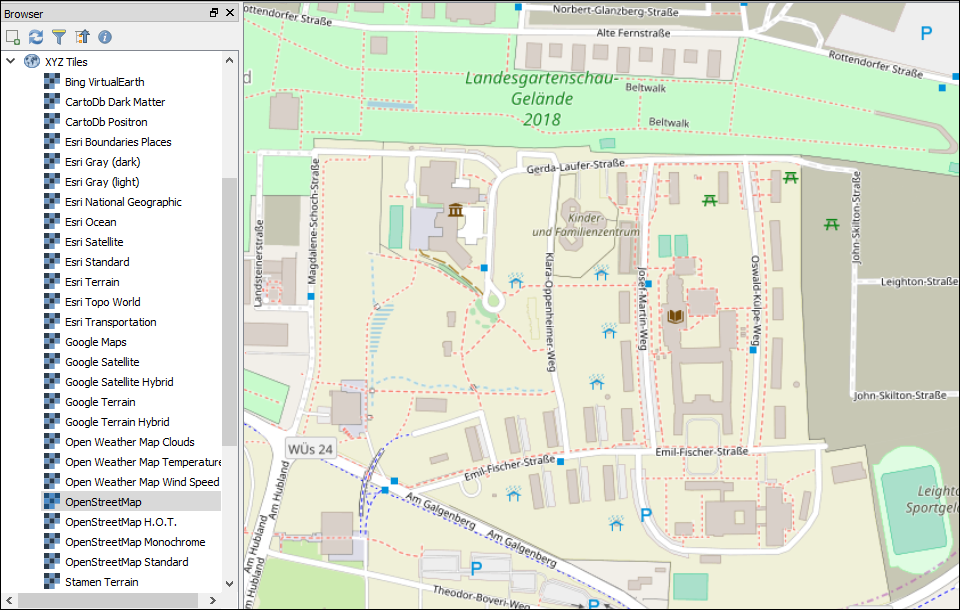
Python Script for QGIS 3.0 Basemaps
The OpenLayer Plugin for QGIS provided all my basemap layers for quite some time now. But since the eagerly awaited launch of QGIS 3.0 earlier this year this plugin became incompatible, leaving you with two options: you either gather and compile all XYZ Tiles...
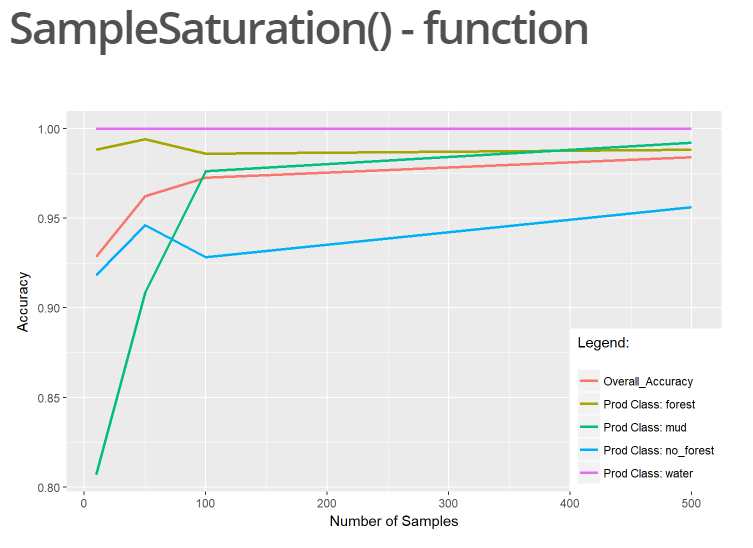
R Package “superClassAnalysis”
For our final project in the course “MB2 – Introduction to Programming and Geostatistics” held by Dr. Martin Wegmann the students were encouraged to explore the possibilities for analyzing and working with remote sensing data in R. In this context I created a package...

Land-Use Change Coded By Color (R Package)
The remote sensing R package LCquickVieweR makes land-use change visible by creating false color images. The methodical approach is a combination of Image Differencing and Multi Temporal Stacking using NDVI. This way change is coded by color. All processing steps are...

To Estimate Biomass Using Light Use Efficiency Model in R
Recently, I published “lue” package in R that is based on the light use efficiency model, which returns the biomass of any crop based on the simple physiological paradigm modelled by Shi et al. (2007). It contains LUE_BIOMASS () which calculates the absorbed...
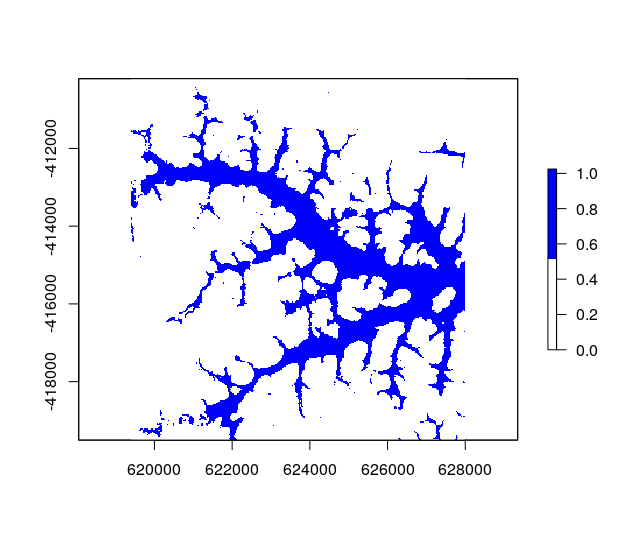
Spectral unmixing in R
In January 2018, I finished the development of the first version of a spectral unmixing function being part of RStoolbox, an R package offering numerous tools for remote sensing analysis written by Benjamin Leutner. The multiple endmember spectral mixture analysis (mesma) function makes it possible to unmix multi- and hyper-spectral imagery by sets of spectral endmember profiles.
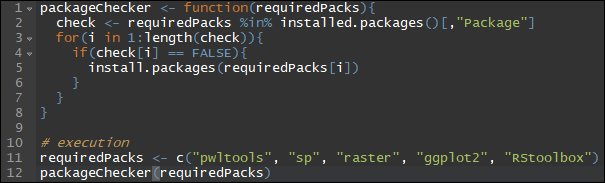
Install every missing R package in one go
Before submitting a project or setting up R on a new system, this function might come in handy. You can put all your favourite or required packages into a single vector and check if they're already installed. If not, packageChecker() will take care of that....
Backslash Converter for R
Every now and then some of us have to work on a campus terminal. And most of the time you want to take the code and data with you. So you do what every student does - you hook up your own usb-drive and change work directories in RStudio. But as soon as you start...
New 2017 EAGLE students welcomed
The new 2017 EAGLE students have been welcomed to Wuerzburg and recently started their studies. Our webpage will be updated as soon as reports and photos about the new students are available.

An EAGLE’s view on the annual congress of the German Society of Photogrammetry and Remote Sensing
In 2017, the German Society of Photogrammetry and Remote Sensing (DGPF) introduced the students’ forum as an additional feature of its annual congress. Thanks to the presentations during this forum, we are now quite well informed of different professional fields in photogrammetry and geodesy. A report.