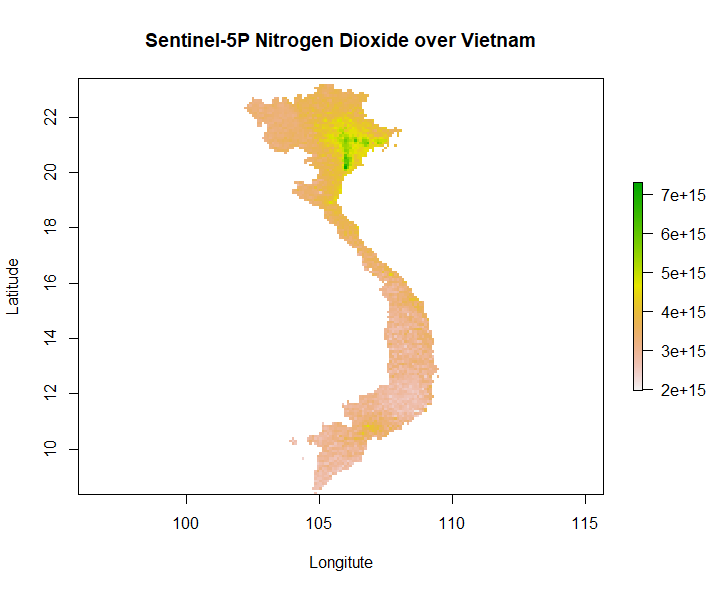
Our second generation EAGLE student Marius Philipp developed the R package S5Processor to automatically convert Sentinel-5P L2 NetCDF files into TIFF files.
The user can define the product which should be turned into a TIFF and also mosaic from a list of file-paths. If no product is defined, the user will be prompted to choose from a list of products. Optionally the spatial resolution of the output imagery, as well as an area of interest to which the imagery will be masked, can be defined. Furthermore, the user can choose whether the pixel values should be either in mol per m² or molecules per cm².
Below an examplary R script:
[code language=”R”]
# Install package from GitHub using devtools.
# For more information visit: https://github.com/MBalthasar/S5Processor
devtools::install_github("MBalthasar/S5Processor")
library(S5Processor)
# Load required packages
library(ncdf4)
library(ggplot2)
library(dismo)
library(maptools)
library(raster)
library(geosphere)
library(rgdal)
library(rgeos)
library(sp)
# Load sample NetCDF files
x1 <- system.file(package = "S5Processor", "extdata", "S5P_NRTI_L2__NO2_1.nc")
x2 <- system.file(package = "S5Processor", "extdata", "S5P_NRTI_L2__NO2_2.nc")
# Create vector from both NetCDF files.
my_files <- c(x1,x2)
# Load sample shapefile including the borders of vietnam
vnm_shp <- raster::shapefile(system.file(package = "S5Processor",
"extdata", "vietnam_borders.shp"))
# Execute function. Defining the spatial resolution in m, product number,
# area of interents and Convert the pixel values to molecules per cm2.
S5P_sample <- S5P_process(input = my_files, my_res = 10000,
product = 39, my_aoi = vnm_shp,
extent_only = FALSE,
apply_scale_factor = T)
[/code]